Antibody knowledge
Recent Breakthroughs in Oligonucleotide Ligation Assay for Disease Detection

Recent breakthroughs in oligonucleotide ligation assay have transformed hiv detection and resistance testing. Researchers now use advanced ligation and assay methods to identify dolutegravir resistance mutations in art-treated patients. The table below shows high detection rates for the assay across diverse variants:
Variant Type | Limit of Detection (copies/mL) | Detection Rate (%) |
|---|---|---|
SARS-CoV-2 | ≥95 |
New ligation techniques increase assay specificity and sensitivity, making diagnostics more reliable. Scientists report improved mutation identification, even at low concentrations. These advances support better art management and guide dolutegravir use in hiv resistance cases.
Improved ligation chemistry gives more consistent assay results.
PCR and qPCR enhance detection of resistance mutations.
Enhanced probe design boosts mutation testing accuracy.
Innovative hybridization-ligation methods enable lower detection limits, which helps identify resistance mutations faster. Healthcare professionals can now rely on oligonucleotide ligation assay for accurate hiv diagnostics and testing of dolutegravir resistance mutations in art regimens.
Key Takeaways
Oligonucleotide ligation assay (OLA) offers highly sensitive and specific detection of genetic mutations, including HIV drug resistance and cancer biomarkers, even at very low levels.
Advanced probe design and ligase selection improve OLA accuracy, reducing false results and enabling detection across diverse viral subtypes and genetic variations.
Integration of PCR and multiplexing allows OLA to detect many mutations simultaneously, speeding up diagnosis and lowering costs for large-scale screening.
OLA is practical and affordable for resource-limited settings, providing rapid, reliable HIV resistance testing that supports timely treatment decisions and better patient outcomes.
Emerging technologies like proximity ligation and next-generation sequencing integration promise to enhance OLA’s sensitivity and personalize disease detection and treatment.
Oligonucleotide Ligation Assay Principles
Probe Design
Probe design plays a vital role in the oligonucleotide ligation assay. Scientists use oligonucleotide probes that match the target sequence exactly at the ligation junction. This perfect match increases specificity and sensitivity. When genetic polymorphisms appear near the ligation site, the assay may fail or give indeterminate results. Researchers found that modifying oligonucleotide probes by changing single nucleotides or several bases near the ligation site restored assay reactivity. For example, in subtype C HIV-1, indeterminate results at certain codons dropped from 66% to 3.3% after probe optimization. Some teams use mixed bases or nonstandard nucleotides to create universal probes. These probes help the oligonucleotide ligation assay work across diverse viral populations.
Tip: Tailored probe design improves both specificity and sensitivity, especially when detecting mutations in genetically diverse samples.
The specificity of the oligonucleotide ligation assay depends on probe-target matching.
Probe optimization reduces indeterminate results and increases detection rates.
Universal probes allow detection of mutations in many subtypes.
DNA Ligase Specificity
DNA ligase is the enzyme that joins oligonucleotide probes during the assay. The specificity of DNA ligase affects the accuracy of mutation detection. T4 DNA ligase can ligate short oligonucleotides quickly, while Tth DNA ligase shows strong inhibition with mismatches. Mismatches at the ligation junction reduce ligation, which increases specificity. Scientists observed low background ligation with mutant probes on wild-type templates, showing high specificity. The oligonucleotide ligation assay detects mutant variants at frequencies as low as 1-3%. The choice of ligase, mismatch position, and oligonucleotide length all influence sensitivity and specificity.
Aspect | Evidence Summary | Impact on OLA Accuracy |
|---|---|---|
Ligase Type | T4 DNA ligase ligates short oligonucleotides rapidly; Tth DNA ligase inhibits ligation with mismatches | Choice of ligase affects mismatch discrimination and speed |
Mismatch Position | Mismatches at or near ligation junction reduce ligation | Enables precise detection of single nucleotide mutations |
Oligonucleotide Length | Pentamers ligated efficiently by T4 DNA ligase; mismatches reduce ligation | High specificity for single nucleotide substitutions |
Assay Format | Solid-phase ligation allows direct detection | Facilitates multiplex detection with high specificity |
Limitations | Only mutations at ligation junction detected | Need for careful assay design to avoid false negatives |
Single Nucleotide Detection
Recent advancements have improved single nucleotide detection in oligonucleotide ligation assay. OLAgen, an open-source software, automates oligonucleotide probe design and PCR assay coupling. This tool aligns sequences around single nucleotide variants and checks for probe dimerization. Assays designed with OLAgen showed 100% sensitivity and specificity in laboratory tests for mutations in Mycobacterium tuberculosis and human genes like JAK2 and BRAF. These assays detect mutant allele frequencies as low as 1% to 5% in wild-type backgrounds. This high sensitivity makes the oligonucleotide ligation assay suitable for clinical diagnostics and research.
OLAgen automates oligonucleotide probe design for single nucleotide detection.
Assays reach 100% sensitivity and specificity for clinically relevant mutations.
Detection of mutant alleles at low frequencies supports early diagnosis.
OLA Technology Advances
PCR-OLA Integration
PCR-OLA integration has changed how scientists detect mutations in clinical samples. This approach combines the power of PCR amplification with the precision of ligation-based assays. Researchers can now detect up to 40 different mutations in a single reaction. The process uses a 15 µl reaction volume with a specific buffer, NAD+, DTT, Triton X-100, OLA probes, PCR product, and Taq ligase. The thermal cycling protocol starts at 95 °C for 1 minute, followed by 32 cycles of 95 °C for 15 seconds and 58 °C for 2 minutes, then cools to 4 °C. This method increases the detection of integrase mutations and improves both sensitivity and specificity.
Aspect | Details |
|---|---|
Multiplexing capability | Up to 40-plex PCR and OLA reactions (e.g., 19-plex SNP assay) |
OLA reaction volume | 15 µl with buffer, NAD+, DTT, Triton X-100, OLA probes, PCR product, Taq ligase |
Thermal cycling protocol | 95 °C for 1 min; 32 cycles of 95 °C for 15 s and 58 °C for 2 min; cooled to 4 °C |
Hybridization | TMAC solution with universal oligonucleotide and FlexMAP microspheres; denature at 95 °C, hybridize at 37 °C |
Fluorescent labeling | Streptavidin-R-phycoerythrin at 37 °C for 40 min; bead sorting and quantitation by Luminex flow cytometry |
Probe design | Two allele-specific probes with unique 24-base FlexMAP tags and one common probe; melting temp ~64 °C |
PCR amplification conditions | 30 µl with AmpliTaq Gold polymerase, dNTPs, primers, MgCl2, 40 ng genomic DNA; 40 cycles |
Plate format | 96-well plates |
Recent improvements in PCR technology, such as advanced thermal cyclers and new reagent chemistries, have made the process faster and more accurate. Microfluidic and digital PCR systems now allow for rapid detection of integrase mutations in many samples at once. Portable PCR devices help bring this technology to clinics and field settings, making detection of mutations more accessible.
Multiplex Detection
Multiplex detection has become a key feature in modern ligation assays. Scientists can now detect thousands of mutations in a single test. The use of T4 RNA Ligase 2 (Rnl2) allows efficient ligation of DNA probes on RNA templates, which helps identify integrase mutations in both DNA and RNA viruses. The cRASL-seq method uses an oligonucleotide-mediated capture step to separate target RNA from excess probes, which increases specificity.
Aspect | Evidence Summary |
|---|---|
Multiplexing Scale | >90% signal retention with 10,000-fold excess irrelevant probes; >5,000 probe sets used; up to 10,000 possible |
Enzyme Innovation | T4 RNA Ligase 2 enables efficient ligation of DNA probes on RNA templates |
Assay Specificity | cRASL-seq capture step enhances specificity |
Pathogen Diversity | Detects fungi, bacteria, DNA and RNA viruses in one assay |
Sensitivity & Quantification | Linear probe read counts; detection limits from ~1.5 to ~150 CFU or PFU/mL |
Sample Multiplexing | DNA barcodes in primers allow high sample throughput and low cost |
Challenges | Index misassignment can reduce specificity; solved by unique dual-indexing |
Limitations | Requires target selection; ~24h turnaround with NGS |
Future Directions | Simultaneous pathogen and host response analysis; point-of-care readouts possible |
Multiplex detection supports the identification of many integrase mutations at once. This capability is important for tracking drug resistance in viruses and for large-scale disease surveillance. The use of DNA barcodes in primers allows scientists to process many samples together, which lowers costs and increases throughput. The main challenge is index misassignment, but unique dual-indexing methods help maintain specificity.
Solid-Phase Protocols
Solid-phase protocols have made high-throughput ligation assays possible. Scientists use controlled-pore glass columns to anchor oligonucleotide strands, which allows for high-yield synthesis. However, this method only supports hundreds of reactions at once. Array-based synthesis methods, such as photochemical or inkjet printing, now allow millions of oligonucleotides to be made in parallel on a single surface. This high level of parallelization supports large-scale detection of integrase mutations.
Solid-phase ligation assays are faster, more sensitive, and more economical than traditional sequencing for detecting drug-resistant mutations. The specificity of the assay depends on the ligase enzyme, which only joins perfectly matched probes. This feature allows for multiplexed detection of many mutations in one test. Solid-phase protocols support efficient and scalable detection of integrase mutations in high-throughput settings.
Note: Software tools like OLAgen help optimize probe design for these advanced assays. OLAgen automates the process, aligns sequences, and checks for probe dimerization. It also designs PCR primers and screens for spurious ligation, which reduces the need for extra lab work. OLAgen-designed assays detect low-frequency integrase mutations with high sensitivity and accuracy, making assay development faster and more reliable.
Applications in Disease Detection
Genetic Disorders
Scientists use advanced detection methods to identify genetic disorders early. The oligonucleotide ligation assay helps detect mutations in genes linked to diseases like cystic fibrosis and sickle cell anemia. This method finds single nucleotide changes that cause these disorders. Researchers use the assay to screen newborns for genetic diseases. Early detection allows doctors to start treatment sooner.
Doctors rely on this technology for population screening. The assay can find rare mutations in large groups. This supports public health programs and helps prevent severe symptoms. The detection of genetic diseases with high sensitivity improves patient outcomes. The assay also works well with clinical samples of hiv-1, showing its flexibility in different settings.
Note: Early detection of genetic mutations helps families make informed decisions about care and treatment.
Cancer Biomarkers
Doctors use the oligonucleotide ligation assay to detect cancer biomarkers. This method finds mutations in genes like JAK2 and BRAF, which play a role in cancer development. The assay detects mutant sequences even when they make up only 0.6% of the sample. This high sensitivity allows for early detection of cancer.
The table below compares the oligonucleotide ligation assay with older molecular diagnostic methods:
Feature/Aspect | Oligonucleotide Ligation Assay (OLA) | Previous Molecular Diagnostic Methods (e.g., Consensus Sequencing, Line Probe Assay) |
|---|---|---|
Sensitivity | Detects mutant sequences as low as 0.6% amid wild-type sequences | Detects predominantly major sequences; minor variants often missed |
Detection Limit | Approximately 19.2 copies per reaction (2.4 × 10^4 copies/ml) | Higher detection limits; less sensitive to low-frequency mutations |
Speed and Throughput | Rapid assay completed within 1 working day; suitable for large-scale screening | Longer turnaround times; less suitable for high-throughput screening |
Cost and Practicality | Economical, easy to interpret visually or via spectrophotometer | More expensive and complex; sequencing requires specialized analysis |
Specificity | High specificity for known point mutations; reduced if nearby mutations exist | Generally high but may miss minor variants or be confounded by mixed populations |
Ability to Detect Mixed Populations | Can detect both wild-type and mutant sequences in mixed samples | Often detects only predominant sequence; minor populations missed |
Limitations | False negatives possible due to probe-target mismatches or additional mutations | Sequencing may fail to detect low-frequency variants; line probe assays have low sensitivity for some mutations |
Comparison with Next-Generation Sequencing (NGS) | Higher sensitivity for low-frequency variants; faster and cheaper | NGS detects novel variants and has high throughput but is costlier, slower, and less sensitive for minor variants |
Doctors use this assay for cancer screening and monitoring. The method supports high-throughput detection of mutations in large patient groups. This helps identify patients who may benefit from targeted therapies. The assay also helps track changes in cancer biomarkers over time.
HIV Resistance
Doctors and researchers use the oligonucleotide ligation assay to detect hiv drug resistance mutations. This method plays a key role in art management. The assay identifies mutations in the hiv-1 integrase gene, as well as nrti and nnrti mutations. These mutations can cause resistance to drugs like dolutegravir, an integrase strand transfer inhibitor.
The assay detects pretreatment hiv drug resistance in patients starting art.
OLA-guided therapy reduces virologic failure in patients with high resistance.
The method detects resistance mutations as well as sequencing methods.
Doctors use the assay to guide art regimen selection and avoid virologic failure.
Studies with 991 hiv-infected participants show the assay works well in real-world settings.
Doctors use OLA-Simple kits for rapid and low-cost detection of resistance mutations. These kits work with common lab equipment and provide results quickly. OLA-Simple detects dolutegravir resistance mutations with 96.5% sensitivity and 100% specificity. The assay works across multiple hiv-1 subtypes, including A, B, C, D, and CRF01_AE. Doctors use the results to make fast clinical decisions.
Tip: OLA-Simple kits help clinics in resource-limited settings perform drug resistance testing and improve patient care.
The oligonucleotide ligation assay, combined with isothermal amplification, labels mutant and wild type hiv-1 sequences. Doctors use a lateral flow ELISA to read the results. This method achieves 100% sensitivity and specificity for the M184V mutation. The assay supports low-cost, decentralized testing in rural hospitals. This helps doctors monitor resistance and make timely decisions.
Doctors use this technology to detect integrase mutations, nrti and nnrti mutations, and dolutegravir resistance. The assay supports genotypic testing and monitoring of art effectiveness. It helps prevent virologic failure and guides changes in therapy. The detection of resistance mutations in clinical samples of hiv-1 improves patient outcomes and supports public health.
Practical Impact in Diagnostics
Resource-Limited Settings
Many clinics in low-income regions face challenges in hiv drug resistance testing. The oligonucleotide ligation assay offers a practical solution for these settings. Clinical studies show that this method detects hiv mutations with high sensitivity and specificity. It identifies minor mutant populations, which helps doctors design effective art regimens. The assay does not need expensive machines or highly trained staff. Health workers in West Africa used this approach to test 122 samples. They found that the assay worked well for early detection of resistance and guided therapy choices.
Attribute | Evidence Summary |
|---|---|
Sensitivity | 98.8% overall sensitivity in detecting HIV-2 drug resistance mutations (Q151M and M184V) |
Specificity | High specificity due to DNA ligase requirements and probe design, with only 1.2% indeterminate results |
Cost | Approximately $1 per sample for two mutations, significantly cheaper than sequencing (~$10 per sample) |
Simplicity & Adaptability | Simple to perform, interpret, and adaptable to new mutations without expensive equipment or expertise |
Clinical Setting | Successfully evaluated with 122 samples from resource-limited countries (West Africa) |
Practical Impact | Enables early detection of drug resistance mutations, guiding therapy in resource-poor settings |
Doctors can use this assay to detect dolutegravir resistance and integrase mutations. This supports better art management and reduces virologic failure.
Workflow Efficiency
The OLA-Simple assay improves workflow in hiv diagnostics. It delivers results in less than four hours, much faster than traditional sequencing, which can take up to three days. Staff with little training can perform the test and still achieve high accuracy. The assay shows about 99.6% specificity and 98.2% sensitivity compared to Sanger sequencing. Clinics can make same-day decisions for art changes, which helps prevent virologic failure. The cost per test is low, and the process does not require complex equipment. This makes routine testing for dolutegravir resistance and integrase mutations more accessible.
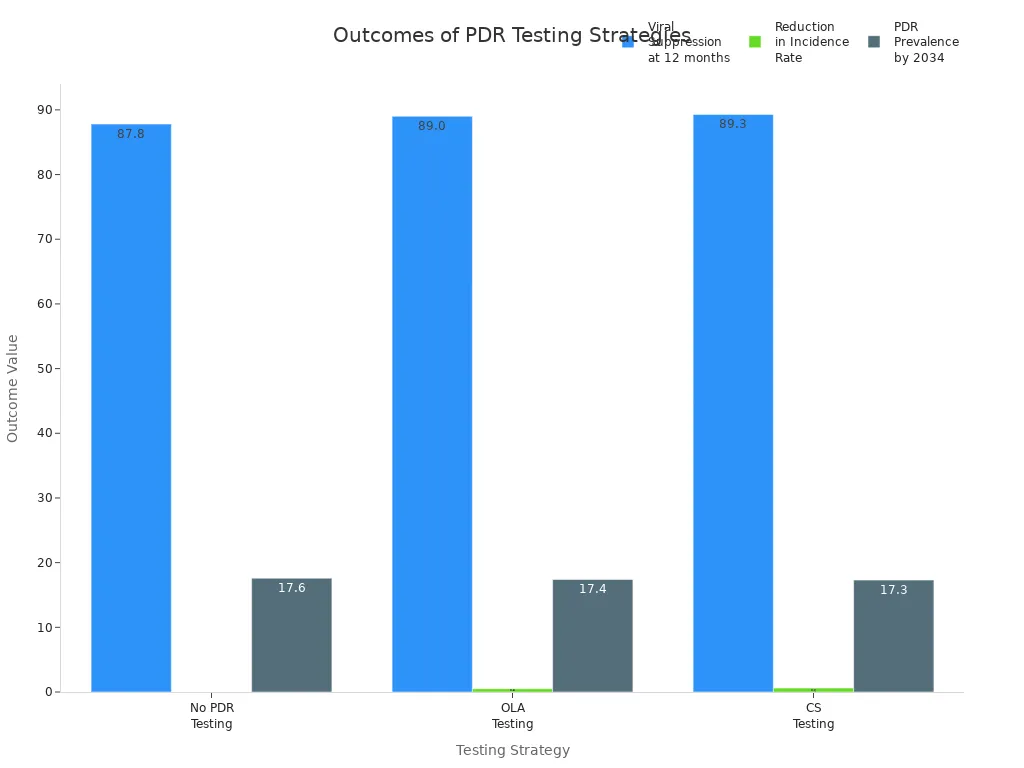
A study in Kenya found that OLA testing improved viral suppression rates but was not cost-effective by national standards. The incremental cost-effectiveness ratio was $10,741 per QALY gained. Despite this, the assay still offers operational benefits for clinics needing rapid detection of hiv mutations.
High-Throughput Screening
High-throughput screening is important for large-scale hiv drug resistance monitoring. The oligonucleotide ligation assay allows labs to test many samples at once. This supports detection of dolutegravir resistance, integrase mutations, and other key changes in hiv. The assay works well for detection of pathogens in mixed infections. Scientists in Thailand used this method to monitor hiv drug resistance in public health programs. The assay detects mutations at levels as low as 3%, while sequencing only finds changes above 20%. This higher sensitivity helps with early detection and better monitoring of art effectiveness.
Public health agencies use OLA kits provided by NIH to support global surveillance. These kits help track dolutegravir resistance and integrase mutations in many countries. The simple design and low cost make the assay suitable for resource-limited settings. Regulatory agencies recognize the value of this approach for improving global health outcomes. The assay supports rapid, accurate, and affordable drug resistance testing, which is vital for controlling hiv and reducing virologic failure worldwide.
Note: The oligonucleotide ligation assay continues to shape the future of hiv diagnostics by making detection, monitoring, and management of resistance mutations more efficient and accessible.
Future Directions
Proximity Ligation
Proximity ligation is changing how scientists detect disease biomarkers. This method uses antibodies attached to oligonucleotides. When both antibodies bind to the same target, the oligonucleotides come close together. A bridging oligonucleotide and DNA ligase then join them. The resulting DNA product is amplified, making even small amounts of target easy to find. Proximity ligation increases sensitivity for detecting integrase mutations and dolutegravir resistance in hiv samples. Researchers can now detect targets at femtomolar levels, which is much lower than traditional tests. Circular proximity ligation (c-PLA) further improves results by reducing background noise and allowing use of low-affinity antibodies. This method works with small sample volumes and can detect multiple biomarkers in plasma, helping doctors diagnose and monitor diseases like hiv and track dolutegravir resistance.
Aspect | Description |
|---|---|
Principle | Antibody-oligonucleotide pairs bind the same target, enabling ligation. |
Amplification | Ligated DNA is amplified by PCR, increasing sensitivity. |
Specificity | Only targets with both probes bound produce a signal. |
Clinical Use | Detects integrase mutations and dolutegravir resistance in hiv samples. |
Next-Gen Sequencing Integration
Oligonucleotide ligation assay is moving toward integration with next-generation sequencing (NGS). Scientists use enzymatic ligation to assemble long l-oligonucleotides from shorter pieces. This supports mirror-image ligase chain reaction and bioconjugation. By combining ligation with NGS, researchers can analyze thousands of integrase mutations and dolutegravir resistance markers in one run. The development of cross-chiral protein ligases and native protein ligases with d-ribonucleotides at the junction allows for more efficient assembly of l-RNA strands. These advances help track hiv mutations and resistance patterns across large populations. The ability to design stable heterochiral complexes also supports new hybridization-based technologies for detecting dolutegravir resistance and integrase mutations.
Scientists develop enzymatic ligation for l-oligonucleotides, making assembly easier.
Native protein ligases with d-ribonucleotides enable ligation of l-RNA strands.
Stable heterochiral complexes improve hybridization for resistance detection.
Long l-oligonucleotides assembled by ligation support NGS and new therapies.
In vitro evolution creates ligases for large-scale production and improved activity.
Personalized Medicine
Personalized medicine uses oligonucleotide ligation assay to match treatments to each patient’s genetic profile. This assay, combined with single-molecule fluorescence, detects even a single nucleotide change in hiv, integrase, or dolutegravir resistance genes. Doctors can identify low-abundance mutations without preamplification. This high sensitivity helps tailor therapy for patients with rare resistance mutations. Accurate detection of integrase mutations and dolutegravir resistance supports better outcomes for people living with hiv. As scientists learn more about how l-oligonucleotides behave in cells, they can design therapies that work better for each person. The future of disease detection will rely on these precise, patient-specific approaches.
Recent breakthroughs in oligonucleotide ligation assay have transformed disease detection. Scientists now detect hiv mutations and resistance with high accuracy and speed. The assay identifies dolutegravir resistance in clinical samples. Researchers use microarray platforms to reach 99.18% diagnostic effectiveness. Multiplex PCR with microarray or Luminex technologies achieves up to 100% sensitivity for respiratory pathogens. Clinics worldwide rely on OLA for hiv resistance testing. The method finds dolutegravir resistance in diverse populations. Doctors use OLA to guide hiv therapy and monitor resistance. The assay supports rapid detection of dolutegravir resistance in resource-limited settings. Global health programs benefit from timely hiv resistance data.
Aspect | |
|---|---|
Application Focus | OLA detects hiv drug resistance mutations, critical for managing hiv globally. |
Advantages | High specificity, cost-effectiveness, and multiplexing support resource-limited settings. |
Technological Advances | OLA with PCR and real-time detection improves sensitivity and speed. |
Practical Impact | Enables timely detection of hiv resistance, supporting effective dolutegravir therapy. |
Ongoing research should address buffer incompatibility and analytical sensitivity. Scientists can further improve dolutegravir resistance detection and expand OLA use for other diseases.
FAQ
What is an oligonucleotide ligation assay (OLA)?
Oligonucleotide ligation assay (OLA) is a laboratory test. Scientists use it to find specific genetic changes. The test joins two DNA pieces only if they match a target sequence. This helps doctors detect mutations linked to diseases.
How does OLA improve disease detection compared to older methods?
OLA finds mutations at very low levels, sometimes as low as 0.6% of a sample. Older methods often miss these rare changes. OLA also works faster and costs less, making it useful for clinics and hospitals.
Can OLA detect multiple mutations at once?
Yes, OLA can find many mutations in one test. Scientists use multiplexing to check for up to 40 different changes in a single reaction. This helps doctors screen for several diseases quickly.
Is OLA useful in places with limited resources?
Doctors in resource-limited clinics use OLA because it needs simple equipment and costs about $1 per sample. Studies in West Africa show OLA detects HIV mutations with 98.8% sensitivity and 1.2% indeterminate results.
What diseases can OLA help diagnose?
Doctors use OLA to detect HIV drug resistance, cancer biomarkers, and genetic disorders like cystic fibrosis. The test works with many types of samples, including blood and saliva.

